\begin{tabular}{lllllllll}
\hline
Summary of computational transaction \tabularnewline
Raw Input & view raw input (R code) \tabularnewline
Raw Output & view raw output of R engine \tabularnewline
Computing time & 1 seconds \tabularnewline
R Server & 'Gwilym Jenkins' @ 72.249.127.135 \tabularnewline
\hline
\end{tabular}
%Source: https://freestatistics.org/blog/index.php?pk=73619&T=0
[TABLE]
[ROW][C]Summary of computational transaction[/C][/ROW]
[ROW][C]Raw Input[/C][C]view raw input (R code) [/C][/ROW]
[ROW][C]Raw Output[/C][C]view raw output of R engine [/C][/ROW]
[ROW][C]Computing time[/C][C]1 seconds[/C][/ROW]
[ROW][C]R Server[/C][C]'Gwilym Jenkins' @ 72.249.127.135[/C][/ROW]
[/TABLE]
Source: https://freestatistics.org/blog/index.php?pk=73619&T=0
If you paste this QR Code into your document, anyone with a smartphone or tablet will be able to scan it and view this table in a browser.
If you paste this QR Code into your document, anyone with a smartphone or tablet will be able to scan it and view this table in a browser.
If you paste this QR Code into your document, anyone with a smartphone or tablet will be able to scan it and view this table in a browser.
If you paste this QR Code into your document, anyone with a smartphone or tablet will be able to scan it and view this table in a browser.
If you paste this QR Code into your document, anyone with a smartphone or tablet will be able to scan it and view this table in a browser.
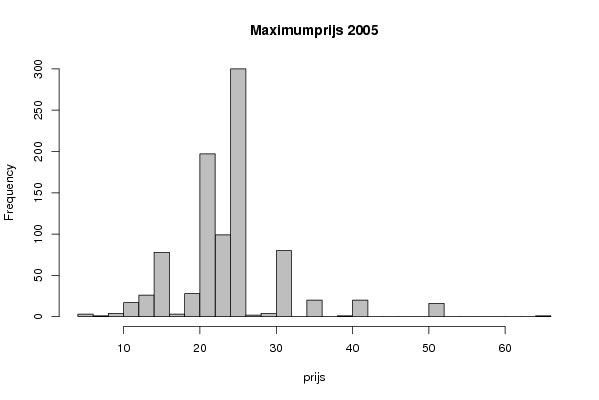
| Frequency Table (Histogram) | | Bins | Midpoint | Abs. Frequency | Rel. Frequency | Cumul. Rel. Freq. | Density | | [4,6[ | 5 | 3 | 0.003333 | 0.003333 | 0.001667 | | [6,8[ | 7 | 1 | 0.001111 | 0.004444 | 0.000556 | | [8,10[ | 9 | 4 | 0.004444 | 0.008889 | 0.002222 | | [10,12[ | 11 | 17 | 0.018889 | 0.027778 | 0.009444 | | [12,14[ | 13 | 26 | 0.028889 | 0.056667 | 0.014444 | | [14,16[ | 15 | 78 | 0.086667 | 0.143333 | 0.043333 | | [16,18[ | 17 | 3 | 0.003333 | 0.146667 | 0.001667 | | [18,20[ | 19 | 28 | 0.031111 | 0.177778 | 0.015556 | | [20,22[ | 21 | 197 | 0.218889 | 0.396667 | 0.109444 | | [22,24[ | 23 | 99 | 0.11 | 0.506667 | 0.055 | | [24,26[ | 25 | 300 | 0.333333 | 0.84 | 0.166667 | | [26,28[ | 27 | 2 | 0.002222 | 0.842222 | 0.001111 | | [28,30[ | 29 | 4 | 0.004444 | 0.846667 | 0.002222 | | [30,32[ | 31 | 80 | 0.088889 | 0.935556 | 0.044444 | | [32,34[ | 33 | 0 | 0 | 0.935556 | 0 | | [34,36[ | 35 | 20 | 0.022222 | 0.957778 | 0.011111 | | [36,38[ | 37 | 0 | 0 | 0.957778 | 0 | | [38,40[ | 39 | 1 | 0.001111 | 0.958889 | 0.000556 | | [40,42[ | 41 | 20 | 0.022222 | 0.981111 | 0.011111 | | [42,44[ | 43 | 0 | 0 | 0.981111 | 0 | | [44,46[ | 45 | 0 | 0 | 0.981111 | 0 | | [46,48[ | 47 | 0 | 0 | 0.981111 | 0 | | [48,50[ | 49 | 0 | 0 | 0.981111 | 0 | | [50,52[ | 51 | 16 | 0.017778 | 0.998889 | 0.008889 | | [52,54[ | 53 | 0 | 0 | 0.998889 | 0 | | [54,56[ | 55 | 0 | 0 | 0.998889 | 0 | | [56,58[ | 57 | 0 | 0 | 0.998889 | 0 | | [58,60[ | 59 | 0 | 0 | 0.998889 | 0 | | [60,62[ | 61 | 0 | 0 | 0.998889 | 0 | | [62,64[ | 63 | 0 | 0 | 0.998889 | 0 | | [64,66] | 65 | 1 | 0.001111 | 1 | 0.000556 |
\begin{tabular}{lllllllll}
\hline
Frequency Table (Histogram) \tabularnewline
Bins & Midpoint & Abs. Frequency & Rel. Frequency & Cumul. Rel. Freq. & Density \tabularnewline
[4,6[ & 5 & 3 & 0.003333 & 0.003333 & 0.001667 \tabularnewline
[6,8[ & 7 & 1 & 0.001111 & 0.004444 & 0.000556 \tabularnewline
[8,10[ & 9 & 4 & 0.004444 & 0.008889 & 0.002222 \tabularnewline
[10,12[ & 11 & 17 & 0.018889 & 0.027778 & 0.009444 \tabularnewline
[12,14[ & 13 & 26 & 0.028889 & 0.056667 & 0.014444 \tabularnewline
[14,16[ & 15 & 78 & 0.086667 & 0.143333 & 0.043333 \tabularnewline
[16,18[ & 17 & 3 & 0.003333 & 0.146667 & 0.001667 \tabularnewline
[18,20[ & 19 & 28 & 0.031111 & 0.177778 & 0.015556 \tabularnewline
[20,22[ & 21 & 197 & 0.218889 & 0.396667 & 0.109444 \tabularnewline
[22,24[ & 23 & 99 & 0.11 & 0.506667 & 0.055 \tabularnewline
[24,26[ & 25 & 300 & 0.333333 & 0.84 & 0.166667 \tabularnewline
[26,28[ & 27 & 2 & 0.002222 & 0.842222 & 0.001111 \tabularnewline
[28,30[ & 29 & 4 & 0.004444 & 0.846667 & 0.002222 \tabularnewline
[30,32[ & 31 & 80 & 0.088889 & 0.935556 & 0.044444 \tabularnewline
[32,34[ & 33 & 0 & 0 & 0.935556 & 0 \tabularnewline
[34,36[ & 35 & 20 & 0.022222 & 0.957778 & 0.011111 \tabularnewline
[36,38[ & 37 & 0 & 0 & 0.957778 & 0 \tabularnewline
[38,40[ & 39 & 1 & 0.001111 & 0.958889 & 0.000556 \tabularnewline
[40,42[ & 41 & 20 & 0.022222 & 0.981111 & 0.011111 \tabularnewline
[42,44[ & 43 & 0 & 0 & 0.981111 & 0 \tabularnewline
[44,46[ & 45 & 0 & 0 & 0.981111 & 0 \tabularnewline
[46,48[ & 47 & 0 & 0 & 0.981111 & 0 \tabularnewline
[48,50[ & 49 & 0 & 0 & 0.981111 & 0 \tabularnewline
[50,52[ & 51 & 16 & 0.017778 & 0.998889 & 0.008889 \tabularnewline
[52,54[ & 53 & 0 & 0 & 0.998889 & 0 \tabularnewline
[54,56[ & 55 & 0 & 0 & 0.998889 & 0 \tabularnewline
[56,58[ & 57 & 0 & 0 & 0.998889 & 0 \tabularnewline
[58,60[ & 59 & 0 & 0 & 0.998889 & 0 \tabularnewline
[60,62[ & 61 & 0 & 0 & 0.998889 & 0 \tabularnewline
[62,64[ & 63 & 0 & 0 & 0.998889 & 0 \tabularnewline
[64,66] & 65 & 1 & 0.001111 & 1 & 0.000556 \tabularnewline
\hline
\end{tabular}
%Source: https://freestatistics.org/blog/index.php?pk=73619&T=1
[TABLE]
[ROW][C]Frequency Table (Histogram)[/C][/ROW]
[ROW][C]Bins[/C][C]Midpoint[/C][C]Abs. Frequency[/C][C]Rel. Frequency[/C][C]Cumul. Rel. Freq.[/C][C]Density[/C][/ROW]
[ROW][C][4,6[[/C][C]5[/C][C]3[/C][C]0.003333[/C][C]0.003333[/C][C]0.001667[/C][/ROW]
[ROW][C][6,8[[/C][C]7[/C][C]1[/C][C]0.001111[/C][C]0.004444[/C][C]0.000556[/C][/ROW]
[ROW][C][8,10[[/C][C]9[/C][C]4[/C][C]0.004444[/C][C]0.008889[/C][C]0.002222[/C][/ROW]
[ROW][C][10,12[[/C][C]11[/C][C]17[/C][C]0.018889[/C][C]0.027778[/C][C]0.009444[/C][/ROW]
[ROW][C][12,14[[/C][C]13[/C][C]26[/C][C]0.028889[/C][C]0.056667[/C][C]0.014444[/C][/ROW]
[ROW][C][14,16[[/C][C]15[/C][C]78[/C][C]0.086667[/C][C]0.143333[/C][C]0.043333[/C][/ROW]
[ROW][C][16,18[[/C][C]17[/C][C]3[/C][C]0.003333[/C][C]0.146667[/C][C]0.001667[/C][/ROW]
[ROW][C][18,20[[/C][C]19[/C][C]28[/C][C]0.031111[/C][C]0.177778[/C][C]0.015556[/C][/ROW]
[ROW][C][20,22[[/C][C]21[/C][C]197[/C][C]0.218889[/C][C]0.396667[/C][C]0.109444[/C][/ROW]
[ROW][C][22,24[[/C][C]23[/C][C]99[/C][C]0.11[/C][C]0.506667[/C][C]0.055[/C][/ROW]
[ROW][C][24,26[[/C][C]25[/C][C]300[/C][C]0.333333[/C][C]0.84[/C][C]0.166667[/C][/ROW]
[ROW][C][26,28[[/C][C]27[/C][C]2[/C][C]0.002222[/C][C]0.842222[/C][C]0.001111[/C][/ROW]
[ROW][C][28,30[[/C][C]29[/C][C]4[/C][C]0.004444[/C][C]0.846667[/C][C]0.002222[/C][/ROW]
[ROW][C][30,32[[/C][C]31[/C][C]80[/C][C]0.088889[/C][C]0.935556[/C][C]0.044444[/C][/ROW]
[ROW][C][32,34[[/C][C]33[/C][C]0[/C][C]0[/C][C]0.935556[/C][C]0[/C][/ROW]
[ROW][C][34,36[[/C][C]35[/C][C]20[/C][C]0.022222[/C][C]0.957778[/C][C]0.011111[/C][/ROW]
[ROW][C][36,38[[/C][C]37[/C][C]0[/C][C]0[/C][C]0.957778[/C][C]0[/C][/ROW]
[ROW][C][38,40[[/C][C]39[/C][C]1[/C][C]0.001111[/C][C]0.958889[/C][C]0.000556[/C][/ROW]
[ROW][C][40,42[[/C][C]41[/C][C]20[/C][C]0.022222[/C][C]0.981111[/C][C]0.011111[/C][/ROW]
[ROW][C][42,44[[/C][C]43[/C][C]0[/C][C]0[/C][C]0.981111[/C][C]0[/C][/ROW]
[ROW][C][44,46[[/C][C]45[/C][C]0[/C][C]0[/C][C]0.981111[/C][C]0[/C][/ROW]
[ROW][C][46,48[[/C][C]47[/C][C]0[/C][C]0[/C][C]0.981111[/C][C]0[/C][/ROW]
[ROW][C][48,50[[/C][C]49[/C][C]0[/C][C]0[/C][C]0.981111[/C][C]0[/C][/ROW]
[ROW][C][50,52[[/C][C]51[/C][C]16[/C][C]0.017778[/C][C]0.998889[/C][C]0.008889[/C][/ROW]
[ROW][C][52,54[[/C][C]53[/C][C]0[/C][C]0[/C][C]0.998889[/C][C]0[/C][/ROW]
[ROW][C][54,56[[/C][C]55[/C][C]0[/C][C]0[/C][C]0.998889[/C][C]0[/C][/ROW]
[ROW][C][56,58[[/C][C]57[/C][C]0[/C][C]0[/C][C]0.998889[/C][C]0[/C][/ROW]
[ROW][C][58,60[[/C][C]59[/C][C]0[/C][C]0[/C][C]0.998889[/C][C]0[/C][/ROW]
[ROW][C][60,62[[/C][C]61[/C][C]0[/C][C]0[/C][C]0.998889[/C][C]0[/C][/ROW]
[ROW][C][62,64[[/C][C]63[/C][C]0[/C][C]0[/C][C]0.998889[/C][C]0[/C][/ROW]
[ROW][C][64,66][/C][C]65[/C][C]1[/C][C]0.001111[/C][C]1[/C][C]0.000556[/C][/ROW]
[/TABLE]
Source: https://freestatistics.org/blog/index.php?pk=73619&T=1
Globally Unique Identifier (entire table): ba.freestatistics.org/blog/index.php?pk=73619&T=1
As an alternative you can also use a QR Code:
The GUIDs for individual cells are displayed in the table below:
| Frequency Table (Histogram) | | Bins | Midpoint | Abs. Frequency | Rel. Frequency | Cumul. Rel. Freq. | Density | | [4,6[ | 5 | 3 | 0.003333 | 0.003333 | 0.001667 | | [6,8[ | 7 | 1 | 0.001111 | 0.004444 | 0.000556 | | [8,10[ | 9 | 4 | 0.004444 | 0.008889 | 0.002222 | | [10,12[ | 11 | 17 | 0.018889 | 0.027778 | 0.009444 | | [12,14[ | 13 | 26 | 0.028889 | 0.056667 | 0.014444 | | [14,16[ | 15 | 78 | 0.086667 | 0.143333 | 0.043333 | | [16,18[ | 17 | 3 | 0.003333 | 0.146667 | 0.001667 | | [18,20[ | 19 | 28 | 0.031111 | 0.177778 | 0.015556 | | [20,22[ | 21 | 197 | 0.218889 | 0.396667 | 0.109444 | | [22,24[ | 23 | 99 | 0.11 | 0.506667 | 0.055 | | [24,26[ | 25 | 300 | 0.333333 | 0.84 | 0.166667 | | [26,28[ | 27 | 2 | 0.002222 | 0.842222 | 0.001111 | | [28,30[ | 29 | 4 | 0.004444 | 0.846667 | 0.002222 | | [30,32[ | 31 | 80 | 0.088889 | 0.935556 | 0.044444 | | [32,34[ | 33 | 0 | 0 | 0.935556 | 0 | | [34,36[ | 35 | 20 | 0.022222 | 0.957778 | 0.011111 | | [36,38[ | 37 | 0 | 0 | 0.957778 | 0 | | [38,40[ | 39 | 1 | 0.001111 | 0.958889 | 0.000556 | | [40,42[ | 41 | 20 | 0.022222 | 0.981111 | 0.011111 | | [42,44[ | 43 | 0 | 0 | 0.981111 | 0 | | [44,46[ | 45 | 0 | 0 | 0.981111 | 0 | | [46,48[ | 47 | 0 | 0 | 0.981111 | 0 | | [48,50[ | 49 | 0 | 0 | 0.981111 | 0 | | [50,52[ | 51 | 16 | 0.017778 | 0.998889 | 0.008889 | | [52,54[ | 53 | 0 | 0 | 0.998889 | 0 | | [54,56[ | 55 | 0 | 0 | 0.998889 | 0 | | [56,58[ | 57 | 0 | 0 | 0.998889 | 0 | | [58,60[ | 59 | 0 | 0 | 0.998889 | 0 | | [60,62[ | 61 | 0 | 0 | 0.998889 | 0 | | [62,64[ | 63 | 0 | 0 | 0.998889 | 0 | | [64,66] | 65 | 1 | 0.001111 | 1 | 0.000556 |
If you paste this QR Code into your document, anyone with a smartphone or tablet will be able to scan it and view this table in a browser.
If you paste this QR Code into your document, anyone with a smartphone or tablet will be able to scan it and view this table in a browser.
If you paste this QR Code into your document, anyone with a smartphone or tablet will be able to scan it and view this table in a browser.
If you paste this QR Code into your document, anyone with a smartphone or tablet will be able to scan it and view this table in a browser.
If you paste this QR Code into your document, anyone with a smartphone or tablet will be able to scan it and view this table in a browser.
|