if (exists('par1')) {
mylist <- list('aston2',
'a-129041896',
'a-129176523',
'a-129097697',
'a-129032827',
'a-129170761',
'a-119117932',
'a-129030638',
'a-129006976',
'a-129030591',
'a-129064998',
'a-109016797',
'a-129020053',
'a-129044901',
'a-119211908',
'a-129176660',
'a-129084642',
'a-129080792',
'a-129051448',
'a-129035622',
'a-129030258',
'a-129174242',
'a-119121928',
'a-129176420',
'a-129018131',
'a-129061469',
'a-129187987',
'a-129004547',
'a-129176877',
'a-129169178',
'a-129073897',
'a-129013387',
'a-129044521',
'a-129175593',
'a-129128533',
'a-129169293',
'a-129170727',
'a-129041656',
'a-129088721',
'a-129035747',
'a-129085155',
'a-129075673',
'a-119120079',
'a-129147648',
'a-129055893',
'a-129022334',
'a-129038346',
'a-129005636',
'a-129033802',
'a-129080954',
'a-129040383',
'a-129078515',
'a-129027638',
'a-129173533',
'a-129055479',
'a-129021740',
'a-129122863',
'a-129025656',
'a-129052559',
'a-119050352',
'a-94085770',
'a-129057772',
'a-129176361',
'a-129026815',
'a-129009047',
'a-119014907',
'a-129077862',
'a-129171908',
'a-129007102',
'a-129043775',
'a-129054715',
'a-129175607',
'a-129062477',
'a-129022703',
'a-129034120',
'a-119159149',
'a-129047337',
'a-129032436',
'a-129022437',
'a-92985278',
'a-129045724',
'a-129047463',
'a-129022725',
'a-129117641',
'a-129066257',
'a-129065238',
'a-129045517',
'a-129046592',
'a-129057875',
'a-129054874',
'a-129043823',
'a-129036582',
'a-129058724',
'a-129025494',
'a-129020732',
'a-129081722',
'a-129060288',
'a-129024187',
'a-129137074',
'a-129031381',
'a-129048459',
'a-2013',
'a-129085801',
'a-129172916',
'a-129043041',
'a-129028473',
'a-129019714',
'a-129178000',
'a-129180508',
'a-129024202',
'a-129055468',
'a-129065836',
'a-129079774',
'a-129035183',
'a-129042435',
'a-129004640',
'a-73406901',
'a-129032090',
'a-129030948',
'a-129015451',
'a-129025070',
'a-129017732',
'a-129043568',
'a-129048286',
'a-129071767',
'a-129083058',
'a-129175250',
'a-096720464',
'a-129028118',
'a-119042162',
'a-119020159',
'a-129066154',
'a-129045610',
'a-129027982',
'a-129038003',
'a-129046008',
'a-129039055',
'a-129072764',
'a-129031635',
'a-129042066',
'a-103291662',
'a-129035530',
'a-129043225',
'a-129170129',
'a-139100709',
'a-129020916',
'a-119052471',
'a-129176925',
'a-129176774',
'a-129177232',
'a-129175412',
'a-129175386',
'a-129025210',
'a-129043247',
'a-129027649',
'a-129054520',
'a-129064677',
'a-129027384',
'a-129168115',
'a-129043546',
'a-129031392',
'a-129032263',
'a-129043812',
'a-129025612',
'a-129027362')
if (length(which(mylist == par1)) > 0) {
par1 <- paste(par1,'_',runif(1),'_',date(),sep='')
} else {
load(file='createtable')
a<-table.start()
a<-table.row.start(a)
a<-table.element(a,'Error',1,TRUE)
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,'The User ID you entered does not exist! ')
a<-table.row.end(a)
a<-table.end(a)
table.save(a,file='mytable1a.tab')
stop('The User ID you entered does not exist!')
}
library(digest)
myran <- runif(1)
if (myran <= 1) {
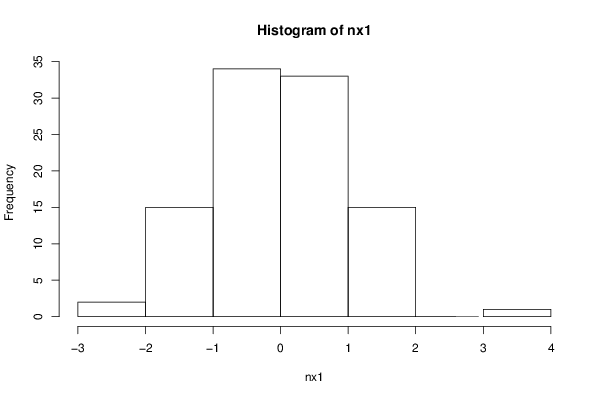
nx1 <- rnorm(100)
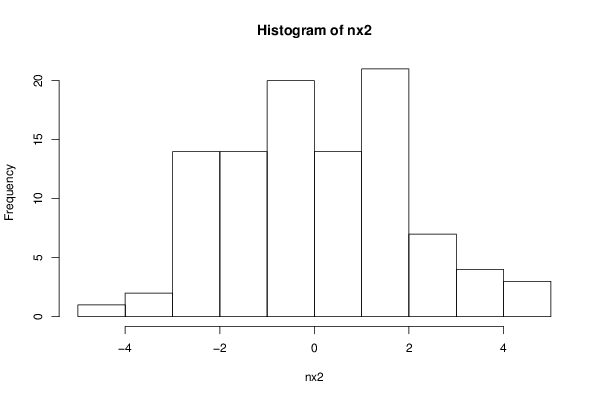
nx2 <- rnorm(100, sd=2.0, mean=0.0)
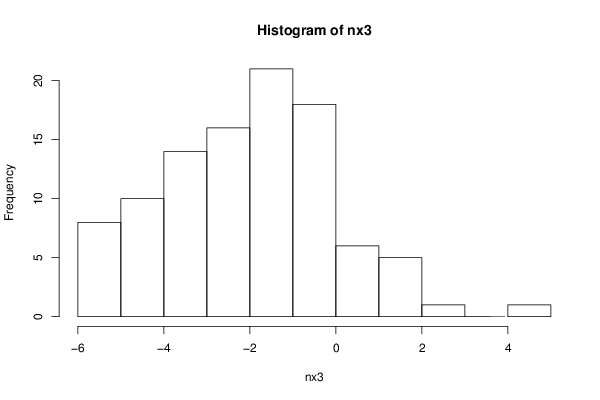
nx3 <- rnorm(100, sd=2.0, mean=-2.0)
answer <- 'nx1'
}
if (myran < 0.66) {
nx2 <- rnorm(100)
nx3 <- rnorm(100, sd=2.0, mean=0.0)
nx1 <- rnorm(100, sd=2.0, mean=-2.0)
answer <- 'nx2'
}
if (myran < 0.33) {
nx3 <- rnorm(100)
nx1 <- rnorm(100, sd=2.0, mean=0.0)
nx2 <- rnorm(100, sd=2.0, mean=-2.0)
answer <- 'nx3'
}
answer <- digest(answer,ser=F)
allmeans<- c(mean(nx1),mean(nx2),mean(nx3))
names(allmeans)<-c('Sample 1', 'Sample 2', 'Sample 3')
allmeans
allsd<- c(sd(nx1), sd(nx2), sd(nx3))
names(allsd)<- c('Sample 1', 'Sample 2', 'Sample 3')
allsd
load(file='createtable')
a<-table.start()
a<-table.row.start(a)
a<-table.element(a,'Identify the sample most similar to a standard normal distribution.',1,TRUE)
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,hyperlink(paste('https://automated.biganalytics.eu/rwasp_May_mock2.wasp?par1=',par1,'&par2=nx1&par3=',answer,'&par0=1',sep=''),'The sample called nx1','',target=''))
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,hyperlink(paste('https://automated.biganalytics.eu/rwasp_May_mock2.wasp?par1=',par1,'&par2=nx2&par3=',answer,'&par0=1',sep=''),'The sample called nx2','',target=''))
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,hyperlink(paste('https://automated.biganalytics.eu/rwasp_May_mock2.wasp?par1=',par1,'&par2=nx3&par3=',answer,'&par0=1',sep=''),'The sample called nx3','',target=''))
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,hyperlink(paste('https://automated.biganalytics.eu/rwasp_May_mock2.wasp?par1=',par1,'&par2=wnga&par3=',answer,'&par0=1',sep=''),'Avoid Guessing Penalty','',target=''))
a<-table.row.end(a)
a<-table.end(a)
table.save(a,file='mytable1.tab')
a<-table.start()
a<-table.row.start(a)
a<-table.element(a,'Means of the three samples',4,TRUE)
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,'Sample',1,TRUE)
a<-table.element(a,'nx1',1,TRUE)
a<-table.element(a,'nx2',1,TRUE)
a<-table.element(a,'nx3',1,TRUE)
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,'Value',1,TRUE)
a<-table.element(a,round(mean(nx1),3))
a<-table.element(a,round(mean(nx2),3))
a<-table.element(a,round(mean(nx3),3))
a<-table.row.end(a)
a<-table.end(a)
table.save(a,file='mytable2.tab')
a<-table.start()
a<-table.row.start(a)
a<-table.element(a,'Standard deviations of the three samples',4,TRUE)
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,'Sample',1,TRUE)
a<-table.element(a,'nx1',1,TRUE)
a<-table.element(a,'nx2',1,TRUE)
a<-table.element(a,'nx3',1,TRUE)
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,'Value',1,TRUE)
a<-table.element(a,round(sd(nx1),3))
a<-table.element(a,round(sd(nx2),3))
a<-table.element(a,round(sd(nx3),3))
a<-table.row.end(a)
a<-table.end(a)
table.save(a,file='mytable3.tab')
bitmap(file='hist1.png')
hist(nx1)
dev.off()
bitmap(file='hist2.png')
hist(nx2)
dev.off()
bitmap(file='hist3.png')
hist(nx3)
dev.off()
bitmap(file='pairs.png')
pairs(x=cbind(nx1, nx2, nx3))
dev.off()
}
|