| Multiple Linear Regression - Estimated Regression Equation |
| K1[t] = + 2.53514 + 0.0442511K2[t] + 0.0677177K3[t] + 0.268577K4[t] + 0.0244293ITH[t] + e[t] |
| Warning: you did not specify the column number of the endogenous series! The first column was selected by default. |
| Multiple Linear Regression - Ordinary Least Squares | |||||
| Variable | Parameter | S.D. | T-STAT H0: parameter = 0 | 2-tail p-value | 1-tail p-value |
| (Intercept) | +2.535 | 0.522 | +4.8570e+00 | 3.131e-06 | 1.566e-06 |
| K2 | +0.04425 | 0.06452 | +6.8580e-01 | 0.494 | 0.247 |
| K3 | +0.06772 | 0.06343 | +1.0680e+00 | 0.2876 | 0.1438 |
| K4 | +0.2686 | 0.06586 | +4.0780e+00 | 7.555e-05 | 3.778e-05 |
| ITH | +0.02443 | 0.02438 | +1.0020e+00 | 0.318 | 0.159 |
| Multiple Linear Regression - Regression Statistics | |
| Multiple R | 0.3646 |
| R-squared | 0.1329 |
| Adjusted R-squared | 0.1083 |
| F-TEST (value) | 5.403 |
| F-TEST (DF numerator) | 4 |
| F-TEST (DF denominator) | 141 |
| p-value | 0.0004458 |
| Multiple Linear Regression - Residual Statistics | |
| Residual Standard Deviation | 0.708 |
| Sum Squared Residuals | 70.67 |
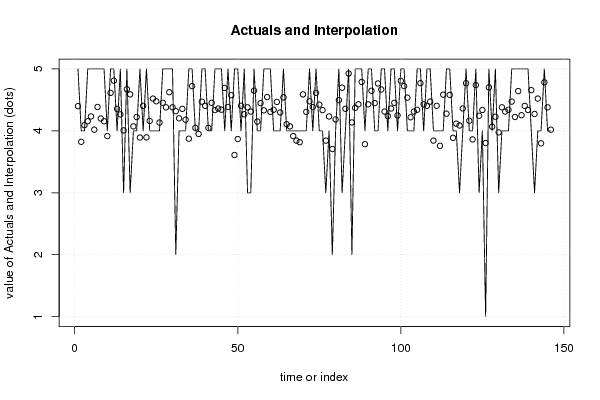
| Multiple Linear Regression - Actuals, Interpolation, and Residuals | |||
| Time or Index | Actuals | Interpolation Forecast | Residuals Prediction Error |
| 1 | 5 | 4.399 | 0.6007 |
| 2 | 4 | 3.824 | 0.1765 |
| 3 | 4 | 4.092 | -0.09208 |
| 4 | 5 | 4.155 | 0.8448 |
| 5 | 5 | 4.233 | 0.7669 |
| 6 | 5 | 4.02 | 0.9802 |
| 7 | 5 | 4.386 | 0.614 |
| 8 | 5 | 4.2 | 0.8002 |
| 9 | 5 | 4.156 | 0.8444 |
| 10 | 4 | 3.917 | 0.08339 |
| 11 | 5 | 4.614 | 0.3864 |
| 12 | 5 | 4.81 | 0.1901 |
| 13 | 4 | 4.356 | -0.356 |
| 14 | 5 | 4.267 | 0.7334 |
| 15 | 3 | 4.009 | -1.009 |
| 16 | 5 | 4.673 | 0.3275 |
| 17 | 3 | 4.59 | -1.59 |
| 18 | 4 | 4.073 | -0.07322 |
| 19 | 4 | 4.224 | -0.2243 |
| 20 | 5 | 3.892 | 1.108 |
| 21 | 4 | 4.405 | -0.4049 |
| 22 | 5 | 3.896 | 1.104 |
| 23 | 4 | 4.16 | -0.1598 |
| 24 | 4 | 4.521 | -0.5215 |
| 25 | 4 | 4.478 | -0.4782 |
| 26 | 4 | 4.135 | -0.1352 |
| 27 | 5 | 4.454 | 0.5462 |
| 28 | 5 | 4.38 | 0.6195 |
| 29 | 5 | 4.625 | 0.3754 |
| 30 | 5 | 4.38 | 0.6195 |
| 31 | 2 | 4.317 | -2.317 |
| 32 | 4 | 4.203 | -0.2031 |
| 33 | 4 | 4.356 | -0.356 |
| 34 | 4 | 4.181 | -0.181 |
| 35 | 5 | 3.873 | 1.127 |
| 36 | 5 | 4.722 | 0.2777 |
| 37 | 4 | 4.049 | -0.04864 |
| 38 | 4 | 3.95 | 0.04989 |
| 39 | 5 | 4.472 | 0.5283 |
| 40 | 5 | 4.404 | 0.5961 |
| 41 | 4 | 4.049 | -0.04879 |
| 42 | 4 | 4.453 | -0.4528 |
| 43 | 5 | 4.336 | 0.6638 |
| 44 | 5 | 4.361 | 0.6393 |
| 45 | 5 | 4.342 | 0.6582 |
| 46 | 4 | 4.693 | -0.6933 |
| 47 | 5 | 4.386 | 0.614 |
| 48 | 4 | 4.576 | -0.5758 |
| 49 | 5 | 3.61 | 1.39 |
| 50 | 5 | 3.868 | 1.132 |
| 51 | 4 | 4.405 | -0.4049 |
| 52 | 5 | 4.269 | 0.7315 |
| 53 | 3 | 4.38 | -1.38 |
| 54 | 3 | 4.311 | -1.311 |
| 55 | 5 | 4.649 | 0.3509 |
| 56 | 4 | 4.151 | -0.1506 |
| 57 | 4 | 4.448 | -0.4482 |
| 58 | 5 | 4.332 | 0.6684 |
| 59 | 5 | 4.546 | 0.4541 |
| 60 | 5 | 4.307 | 0.6928 |
| 61 | 4 | 4.341 | -0.3408 |
| 62 | 4 | 4.467 | -0.4671 |
| 63 | 4 | 4.298 | -0.2975 |
| 64 | 5 | 4.541 | 0.4587 |
| 65 | 4 | 4.107 | -0.1073 |
| 66 | 4 | 4.072 | -0.07225 |
| 67 | 4 | 3.917 | 0.08339 |
| 68 | 4 | 3.842 | 0.1576 |
| 69 | 4 | 3.818 | 0.1821 |
| 70 | 4 | 4.589 | -0.5892 |
| 71 | 4 | 4.307 | -0.3072 |
| 72 | 5 | 4.477 | 0.5228 |
| 73 | 4 | 4.385 | -0.3851 |
| 74 | 5 | 4.614 | 0.3864 |
| 75 | 4 | 4.424 | -0.4238 |
| 76 | 4 | 4.335 | -0.3353 |
| 77 | 3 | 3.842 | -0.8424 |
| 78 | 4 | 4.233 | -0.2331 |
| 79 | 2 | 3.707 | -1.707 |
| 80 | 4 | 4.185 | -0.1852 |
| 81 | 5 | 4.497 | 0.5029 |
| 82 | 3 | 4.698 | -1.698 |
| 83 | 4 | 4.356 | -0.356 |
| 84 | 5 | 4.926 | 0.07354 |
| 85 | 2 | 4.136 | -2.136 |
| 86 | 5 | 4.372 | 0.6284 |
| 87 | 5 | 4.429 | 0.5707 |
| 88 | 5 | 4.79 | 0.2099 |
| 89 | 4 | 3.785 | 0.2152 |
| 90 | 5 | 4.428 | 0.5716 |
| 91 | 5 | 4.648 | 0.3519 |
| 92 | 4 | 4.448 | -0.4482 |
| 93 | 4 | 4.766 | -0.7656 |
| 94 | 5 | 4.668 | 0.3321 |
| 95 | 5 | 4.312 | 0.6882 |
| 96 | 4 | 4.239 | -0.2385 |
| 97 | 5 | 4.361 | 0.6393 |
| 98 | 5 | 4.453 | 0.5472 |
| 99 | 4 | 4.249 | -0.2487 |
| 100 | 5 | 4.809 | 0.1911 |
| 101 | 5 | 4.723 | 0.2767 |
| 102 | 4 | 4.537 | -0.5371 |
| 103 | 4 | 4.22 | -0.2196 |
| 104 | 4 | 4.312 | -0.3118 |
| 105 | 5 | 4.341 | 0.6592 |
| 106 | 5 | 4.77 | 0.2298 |
| 107 | 4 | 4.428 | -0.4284 |
| 108 | 5 | 4.405 | 0.5951 |
| 109 | 5 | 4.473 | 0.5274 |
| 110 | 4 | 3.842 | 0.1576 |
| 111 | 4 | 4.405 | -0.4049 |
| 112 | 4 | 3.759 | 0.2406 |
| 113 | 4 | 4.585 | -0.5846 |
| 114 | 5 | 4.277 | 0.7227 |
| 115 | 5 | 4.58 | 0.4196 |
| 116 | 4 | 3.888 | 0.1124 |
| 117 | 4 | 4.117 | -0.1165 |
| 118 | 3 | 4.087 | -1.087 |
| 119 | 4 | 4.362 | -0.3616 |
| 120 | 5 | 4.77 | 0.2298 |
| 121 | 4 | 4.164 | -0.1644 |
| 122 | 4 | 3.862 | 0.1378 |
| 123 | 5 | 4.74 | 0.2598 |
| 124 | 3 | 4.244 | -1.244 |
| 125 | 4 | 4.337 | -0.3372 |
| 126 | 1 | 3.804 | -2.804 |
| 127 | 5 | 4.703 | 0.2975 |
| 128 | 4 | 4.069 | -0.06861 |
| 129 | 5 | 4.228 | 0.7715 |
| 130 | 3 | 3.975 | -0.9755 |
| 131 | 4 | 4.38 | -0.3805 |
| 132 | 4 | 4.312 | -0.3118 |
| 133 | 4 | 4.341 | -0.3408 |
| 134 | 5 | 4.473 | 0.5274 |
| 135 | 5 | 4.224 | 0.7757 |
| 136 | 5 | 4.643 | 0.3575 |
| 137 | 5 | 4.253 | 0.7471 |
| 138 | 5 | 4.403 | 0.597 |
| 139 | 5 | 4.337 | 0.6628 |
| 140 | 4 | 4.657 | -0.6573 |
| 141 | 3 | 4.273 | -1.273 |
| 142 | 4 | 4.521 | -0.5215 |
| 143 | 4 | 3.799 | 0.2009 |
| 144 | 5 | 4.784 | 0.2155 |
| 145 | 4 | 4.38 | -0.3805 |
| 146 | 4 | 4.019 | -0.01879 |
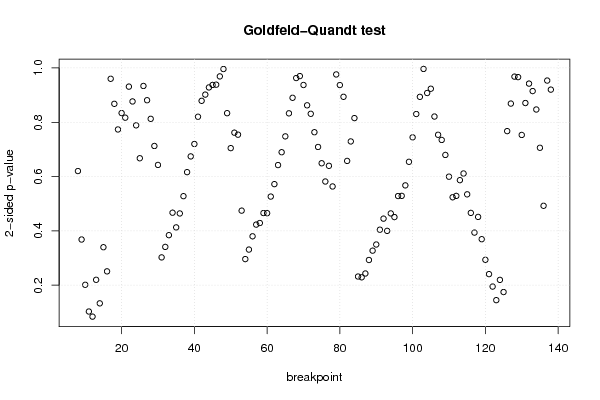
| Goldfeld-Quandt test for Heteroskedasticity | |||
| p-values | Alternative Hypothesis | ||
| breakpoint index | greater | 2-sided | less |
| 8 | 0.3102 | 0.6204 | 0.6898 |
| 9 | 0.1841 | 0.3682 | 0.8159 |
| 10 | 0.1007 | 0.2014 | 0.8993 |
| 11 | 0.05149 | 0.103 | 0.9485 |
| 12 | 0.04201 | 0.08402 | 0.958 |
| 13 | 0.1099 | 0.2198 | 0.8901 |
| 14 | 0.06654 | 0.1331 | 0.9335 |
| 15 | 0.1699 | 0.3397 | 0.8301 |
| 16 | 0.1255 | 0.2509 | 0.8745 |
| 17 | 0.5199 | 0.9603 | 0.4801 |
| 18 | 0.434 | 0.868 | 0.566 |
| 19 | 0.3867 | 0.7735 | 0.6133 |
| 20 | 0.4169 | 0.8339 | 0.5831 |
| 21 | 0.4084 | 0.8169 | 0.5916 |
| 22 | 0.4657 | 0.9313 | 0.5343 |
| 23 | 0.4384 | 0.8767 | 0.5616 |
| 24 | 0.3944 | 0.7888 | 0.6056 |
| 25 | 0.3338 | 0.6675 | 0.6662 |
| 26 | 0.4669 | 0.9337 | 0.5331 |
| 27 | 0.4406 | 0.8812 | 0.5594 |
| 28 | 0.4064 | 0.8127 | 0.5936 |
| 29 | 0.3564 | 0.7127 | 0.6436 |
| 30 | 0.3214 | 0.6428 | 0.6786 |
| 31 | 0.8488 | 0.3023 | 0.1512 |
| 32 | 0.8295 | 0.3409 | 0.1705 |
| 33 | 0.8079 | 0.3842 | 0.1921 |
| 34 | 0.7666 | 0.4669 | 0.2334 |
| 35 | 0.7936 | 0.4128 | 0.2064 |
| 36 | 0.7679 | 0.4642 | 0.2321 |
| 37 | 0.7361 | 0.5278 | 0.2639 |
| 38 | 0.6918 | 0.6163 | 0.3082 |
| 39 | 0.6628 | 0.6743 | 0.3372 |
| 40 | 0.64 | 0.72 | 0.36 |
| 41 | 0.5898 | 0.8204 | 0.4102 |
| 42 | 0.5605 | 0.879 | 0.4395 |
| 43 | 0.549 | 0.902 | 0.451 |
| 44 | 0.5358 | 0.9283 | 0.4642 |
| 45 | 0.5312 | 0.9376 | 0.4688 |
| 46 | 0.5308 | 0.9385 | 0.4692 |
| 47 | 0.5155 | 0.969 | 0.4845 |
| 48 | 0.4981 | 0.9962 | 0.5019 |
| 49 | 0.5833 | 0.8335 | 0.4167 |
| 50 | 0.6476 | 0.7048 | 0.3524 |
| 51 | 0.6192 | 0.7617 | 0.3808 |
| 52 | 0.6229 | 0.7543 | 0.3771 |
| 53 | 0.7628 | 0.4745 | 0.2372 |
| 54 | 0.852 | 0.296 | 0.148 |
| 55 | 0.8345 | 0.3311 | 0.1655 |
| 56 | 0.81 | 0.38 | 0.19 |
| 57 | 0.7885 | 0.423 | 0.2115 |
| 58 | 0.7855 | 0.4289 | 0.2145 |
| 59 | 0.7671 | 0.4657 | 0.2329 |
| 60 | 0.7674 | 0.4652 | 0.2326 |
| 61 | 0.7368 | 0.5263 | 0.2632 |
| 62 | 0.7139 | 0.5721 | 0.2861 |
| 63 | 0.6788 | 0.6424 | 0.3212 |
| 64 | 0.6552 | 0.6897 | 0.3448 |
| 65 | 0.626 | 0.7481 | 0.374 |
| 66 | 0.5835 | 0.833 | 0.4165 |
| 67 | 0.5549 | 0.8902 | 0.4451 |
| 68 | 0.5185 | 0.963 | 0.4815 |
| 69 | 0.4851 | 0.9702 | 0.5149 |
| 70 | 0.4687 | 0.9374 | 0.5313 |
| 71 | 0.4313 | 0.8625 | 0.5687 |
| 72 | 0.4157 | 0.8313 | 0.5843 |
| 73 | 0.3817 | 0.7634 | 0.6183 |
| 74 | 0.3545 | 0.709 | 0.6455 |
| 75 | 0.3245 | 0.6491 | 0.6755 |
| 76 | 0.2909 | 0.5818 | 0.7091 |
| 77 | 0.3198 | 0.6396 | 0.6802 |
| 78 | 0.2818 | 0.5636 | 0.7182 |
| 79 | 0.5119 | 0.9762 | 0.4881 |
| 80 | 0.4686 | 0.9372 | 0.5314 |
| 81 | 0.4469 | 0.8939 | 0.5531 |
| 82 | 0.6712 | 0.6576 | 0.3288 |
| 83 | 0.6353 | 0.7294 | 0.3647 |
| 84 | 0.5923 | 0.8154 | 0.4077 |
| 85 | 0.8841 | 0.2319 | 0.1159 |
| 86 | 0.8855 | 0.229 | 0.1145 |
| 87 | 0.8785 | 0.243 | 0.1215 |
| 88 | 0.8536 | 0.2927 | 0.1464 |
| 89 | 0.8365 | 0.3271 | 0.1635 |
| 90 | 0.8251 | 0.3497 | 0.1749 |
| 91 | 0.7979 | 0.4043 | 0.2021 |
| 92 | 0.7775 | 0.445 | 0.2225 |
| 93 | 0.8 | 0.4 | 0.2 |
| 94 | 0.7679 | 0.4643 | 0.2321 |
| 95 | 0.7746 | 0.4508 | 0.2254 |
| 96 | 0.7358 | 0.5283 | 0.2642 |
| 97 | 0.7355 | 0.5289 | 0.2645 |
| 98 | 0.7162 | 0.5676 | 0.2838 |
| 99 | 0.6728 | 0.6543 | 0.3272 |
| 100 | 0.6278 | 0.7445 | 0.3722 |
| 101 | 0.5848 | 0.8304 | 0.4152 |
| 102 | 0.5532 | 0.8937 | 0.4468 |
| 103 | 0.5018 | 0.9964 | 0.4982 |
| 104 | 0.4539 | 0.9078 | 0.5461 |
| 105 | 0.4618 | 0.9235 | 0.5382 |
| 106 | 0.4106 | 0.8211 | 0.5894 |
| 107 | 0.377 | 0.754 | 0.623 |
| 108 | 0.3675 | 0.7349 | 0.6325 |
| 109 | 0.3398 | 0.6797 | 0.6602 |
| 110 | 0.2998 | 0.5995 | 0.7002 |
| 111 | 0.2619 | 0.5238 | 0.7381 |
| 112 | 0.2644 | 0.5288 | 0.7356 |
| 113 | 0.2934 | 0.5868 | 0.7066 |
| 114 | 0.3056 | 0.6113 | 0.6944 |
| 115 | 0.2673 | 0.5346 | 0.7327 |
| 116 | 0.2331 | 0.4662 | 0.7669 |
| 117 | 0.1968 | 0.3936 | 0.8032 |
| 118 | 0.2257 | 0.4513 | 0.7743 |
| 119 | 0.1846 | 0.3693 | 0.8154 |
| 120 | 0.1467 | 0.2935 | 0.8533 |
| 121 | 0.1203 | 0.2407 | 0.8797 |
| 122 | 0.0974 | 0.1948 | 0.9026 |
| 123 | 0.07241 | 0.1448 | 0.9276 |
| 124 | 0.1097 | 0.2194 | 0.8903 |
| 125 | 0.08734 | 0.1747 | 0.9127 |
| 126 | 0.6163 | 0.7675 | 0.3837 |
| 127 | 0.5656 | 0.8689 | 0.4344 |
| 128 | 0.484 | 0.968 | 0.516 |
| 129 | 0.5168 | 0.9664 | 0.4832 |
| 130 | 0.6234 | 0.7533 | 0.3766 |
| 131 | 0.5645 | 0.871 | 0.4355 |
| 132 | 0.5288 | 0.9424 | 0.4712 |
| 133 | 0.4575 | 0.9149 | 0.5425 |
| 134 | 0.4235 | 0.8471 | 0.5765 |
| 135 | 0.353 | 0.7061 | 0.647 |
| 136 | 0.2462 | 0.4924 | 0.7538 |
| 137 | 0.5232 | 0.9537 | 0.4768 |
| 138 | 0.4601 | 0.9202 | 0.5399 |
| Meta Analysis of Goldfeld-Quandt test for Heteroskedasticity | |||
| Description | # significant tests | % significant tests | OK/NOK |
| 1% type I error level | 0 | 0 | OK |
| 5% type I error level | 0 | 0 | OK |
| 10% type I error level | 1 | 0.00763359 | OK |
| Ramsey RESET F-Test for powers (2 and 3) of fitted values |
> reset_test_fitted RESET test data: mylm RESET = 0.086865, df1 = 2, df2 = 139, p-value = 0.9169 |
| Ramsey RESET F-Test for powers (2 and 3) of regressors |
> reset_test_regressors RESET test data: mylm RESET = 1.4181, df1 = 8, df2 = 133, p-value = 0.1945 |
| Ramsey RESET F-Test for powers (2 and 3) of principal components |
> reset_test_principal_components RESET test data: mylm RESET = 1.6438, df1 = 2, df2 = 139, p-value = 0.197 |
| Variance Inflation Factors (Multicollinearity) |
> vif
K2 K3 K4 ITH
1.044557 1.060569 1.021999 1.010675
|