| Multiple Linear Regression - Estimated Regression Equation |
| TVDC1[t] = -2.45041e-14 -1TVDC2[t] -1TVDC3[t] -1TVDC4[t] + 1TVDCSUM[t] + e[t] |
| Multiple Linear Regression - Ordinary Least Squares | |||||
| Variable | Parameter | S.D. | T-STAT H0: parameter = 0 | 2-tail p-value | 1-tail p-value |
| (Intercept) | -2.45e-14 | 7.477e-15 | -3.2770e+00 | 0.001451 | 0.0007255 |
| TVDC2 | -1 | 1.988e-15 | -5.0300e+14 | 0 | 0 |
| TVDC3 | -1 | 2.156e-15 | -4.6370e+14 | 0 | 0 |
| TVDC4 | -1 | 1.637e-15 | -6.1100e+14 | 0 | 0 |
| TVDCSUM | +1 | 9.819e-16 | +1.0180e+15 | 0 | 0 |
| Multiple Linear Regression - Regression Statistics | |
| Multiple R | 1 |
| R-squared | 1 |
| Adjusted R-squared | 1 |
| F-TEST (value) | 3.508e+29 |
| F-TEST (DF numerator) | 4 |
| F-TEST (DF denominator) | 98 |
| p-value | 0 |
| Multiple Linear Regression - Residual Statistics | |
| Residual Standard Deviation | 8.399e-15 |
| Sum Squared Residuals | 6.913e-27 |
| Menu of Residual Diagnostics | |
| Description | Link |
| Histogram | Compute |
| Central Tendency | Compute |
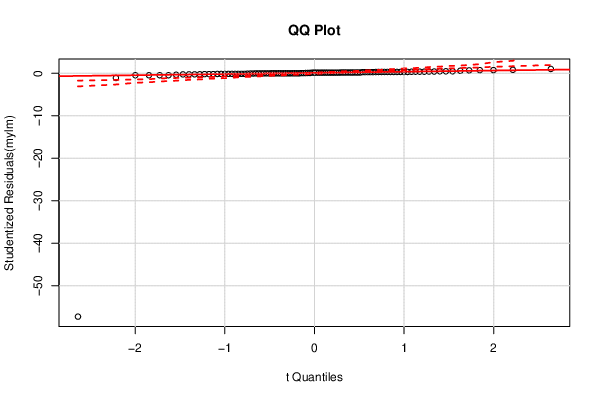
| QQ Plot | Compute |
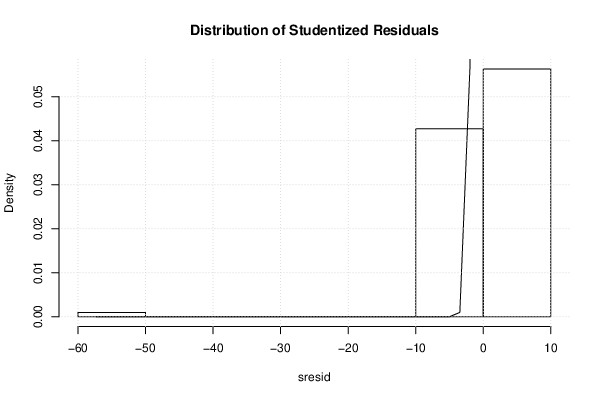
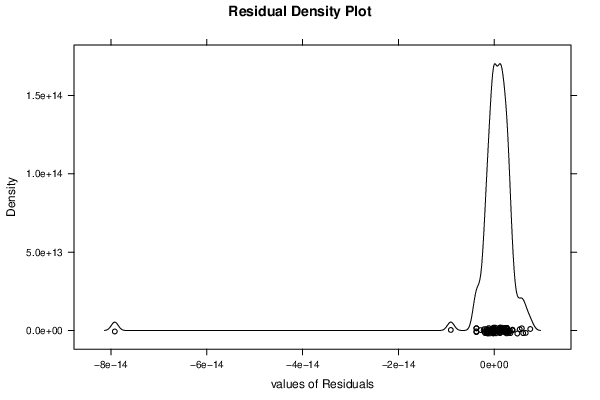
| Kernel Density Plot | Compute |
| Skewness/Kurtosis Test | Compute |
| Skewness-Kurtosis Plot | Compute |
| Harrell-Davis Plot | Compute |
| Bootstrap Plot -- Central Tendency | Compute |
| Blocked Bootstrap Plot -- Central Tendency | Compute |
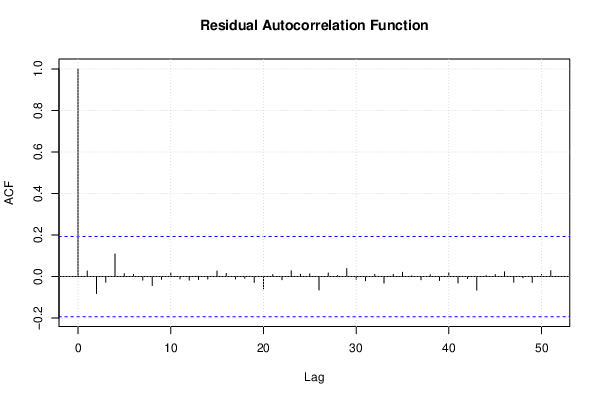
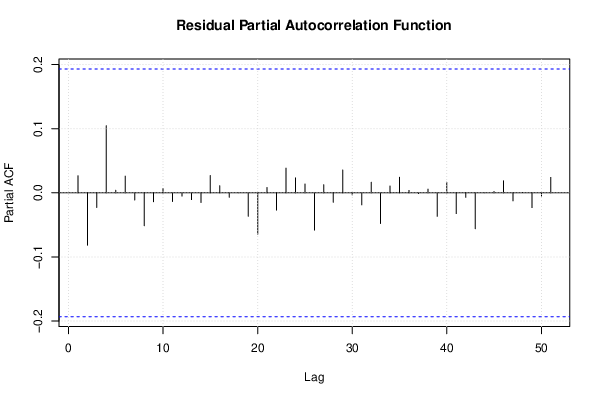
| (Partial) Autocorrelation Plot | Compute |
| Spectral Analysis | Compute |
| Tukey lambda PPCC Plot | Compute |
| Box-Cox Normality Plot | Compute |
| Summary Statistics | Compute |
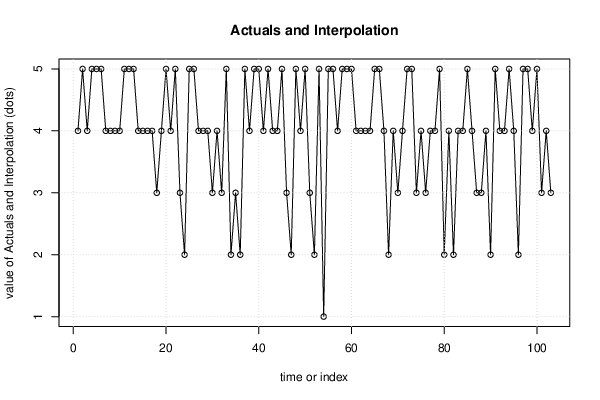
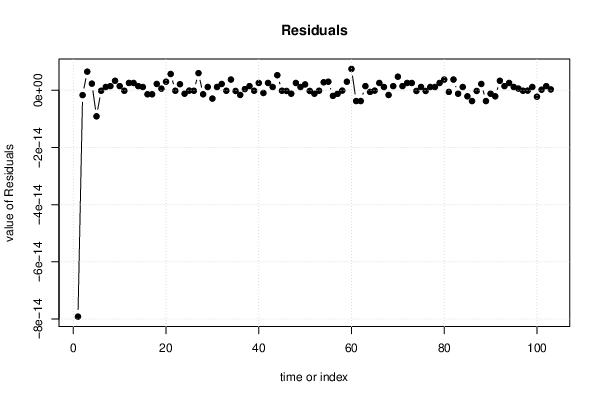
| Multiple Linear Regression - Actuals, Interpolation, and Residuals | |||
| Time or Index | Actuals | Interpolation Forecast | Residuals Prediction Error |
| 1 | 4 | 4 | -7.919e-14 |
| 2 | 5 | 5 | -1.672e-15 |
| 3 | 4 | 4 | 6.541e-15 |
| 4 | 5 | 5 | 2.349e-15 |
| 5 | 5 | 5 | -9.087e-15 |
| 6 | 5 | 5 | -1.413e-16 |
| 7 | 4 | 4 | 1.167e-15 |
| 8 | 4 | 4 | 1.476e-15 |
| 9 | 4 | 4 | 3.344e-15 |
| 10 | 4 | 4 | 1.476e-15 |
| 11 | 5 | 5 | -1.413e-16 |
| 12 | 5 | 5 | 2.587e-15 |
| 13 | 5 | 5 | 2.587e-15 |
| 14 | 4 | 4 | 1.476e-15 |
| 15 | 4 | 4 | 1.167e-15 |
| 16 | 4 | 4 | -1.374e-15 |
| 17 | 4 | 4 | -1.374e-15 |
| 18 | 3 | 3 | 2.24e-15 |
| 19 | 4 | 4 | 5.882e-16 |
| 20 | 5 | 5 | 3.02e-15 |
| 21 | 4 | 4 | 5.723e-15 |
| 22 | 5 | 5 | -1.413e-16 |
| 23 | 3 | 3 | 2.157e-15 |
| 24 | 2 | 2 | -1.177e-15 |
| 25 | 5 | 5 | -9.673e-17 |
| 26 | 5 | 5 | -1.413e-16 |
| 27 | 4 | 4 | 6.017e-15 |
| 28 | 4 | 4 | -1.374e-15 |
| 29 | 4 | 4 | 1.167e-15 |
| 30 | 3 | 3 | -2.884e-15 |
| 31 | 4 | 4 | 1.167e-15 |
| 32 | 3 | 3 | 2.24e-15 |
| 33 | 5 | 5 | -1.413e-16 |
| 34 | 2 | 2 | 3.774e-15 |
| 35 | 3 | 3 | -2.108e-16 |
| 36 | 2 | 2 | -1.61e-15 |
| 37 | 5 | 5 | 4.309e-16 |
| 38 | 4 | 4 | 1.476e-15 |
| 39 | 5 | 5 | -1.413e-16 |
| 40 | 5 | 5 | 2.587e-15 |
| 41 | 4 | 4 | -9.126e-16 |
| 42 | 5 | 5 | 2.587e-15 |
| 43 | 4 | 4 | 1.167e-15 |
| 44 | 4 | 4 | 5.289e-15 |
| 45 | 5 | 5 | -1.413e-16 |
| 46 | 3 | 3 | -2.108e-16 |
| 47 | 2 | 2 | -1.177e-15 |
| 48 | 5 | 5 | 2.587e-15 |
| 49 | 4 | 4 | 1.167e-15 |
| 50 | 5 | 5 | 2.07e-15 |
| 51 | 3 | 3 | -2.108e-16 |
| 52 | 2 | 2 | -1.188e-15 |
| 53 | 5 | 5 | -1.413e-16 |
| 54 | 1 | 1 | 2.82e-15 |
| 55 | 5 | 5 | 3.02e-15 |
| 56 | 5 | 5 | -1.931e-15 |
| 57 | 4 | 4 | -1.218e-15 |
| 58 | 5 | 5 | -9.673e-17 |
| 59 | 5 | 5 | 3.02e-15 |
| 60 | 5 | 5 | 7.489e-15 |
| 61 | 4 | 4 | -3.752e-15 |
| 62 | 4 | 4 | -3.752e-15 |
| 63 | 4 | 4 | 1.476e-15 |
| 64 | 4 | 4 | -5.459e-16 |
| 65 | 5 | 5 | -9.673e-17 |
| 66 | 5 | 5 | 2.587e-15 |
| 67 | 4 | 4 | 1.167e-15 |
| 68 | 2 | 2 | -1.61e-15 |
| 69 | 4 | 4 | 1.476e-15 |
| 70 | 3 | 3 | 4.785e-15 |
| 71 | 4 | 4 | 1.476e-15 |
| 72 | 5 | 5 | 2.587e-15 |
| 73 | 5 | 5 | 2.587e-15 |
| 74 | 3 | 3 | -2.108e-16 |
| 75 | 4 | 4 | 1.167e-15 |
| 76 | 3 | 3 | -2.108e-16 |
| 77 | 4 | 4 | 1.167e-15 |
| 78 | 4 | 4 | 1.167e-15 |
| 79 | 5 | 5 | 2.587e-15 |
| 80 | 2 | 2 | 3.774e-15 |
| 81 | 4 | 4 | -5.459e-16 |
| 82 | 2 | 2 | 3.774e-15 |
| 83 | 4 | 4 | -1.174e-15 |
| 84 | 4 | 4 | 1.167e-15 |
| 85 | 5 | 5 | -2.086e-15 |
| 86 | 4 | 4 | -3.752e-15 |
| 87 | 3 | 3 | -2.108e-16 |
| 88 | 3 | 3 | 2.24e-15 |
| 89 | 4 | 4 | -3.752e-15 |
| 90 | 2 | 2 | -1.177e-15 |
| 91 | 5 | 5 | -2.086e-15 |
| 92 | 4 | 4 | 3.344e-15 |
| 93 | 4 | 4 | 1.476e-15 |
| 94 | 5 | 5 | 2.587e-15 |
| 95 | 4 | 4 | 1.167e-15 |
| 96 | 2 | 2 | 6.19e-16 |
| 97 | 5 | 5 | -1.413e-16 |
| 98 | 5 | 5 | -9.673e-17 |
| 99 | 4 | 4 | 1.167e-15 |
| 100 | 5 | 5 | -2.221e-15 |
| 101 | 3 | 3 | 2.223e-16 |
| 102 | 4 | 4 | 1.476e-15 |
| 103 | 3 | 3 | 3.224e-16 |
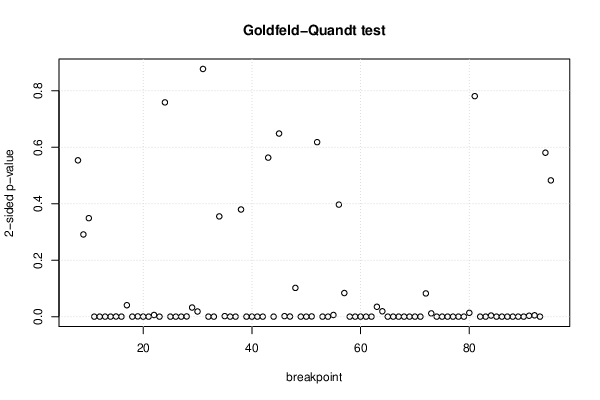
| Goldfeld-Quandt test for Heteroskedasticity | |||
| p-values | Alternative Hypothesis | ||
| breakpoint index | greater | 2-sided | less |
| 8 | 0.277 | 0.5539 | 0.723 |
| 9 | 0.1456 | 0.2912 | 0.8544 |
| 10 | 0.8255 | 0.349 | 0.1745 |
| 11 | 3.204e-06 | 6.409e-06 | 1 |
| 12 | 2.51e-06 | 5.02e-06 | 1 |
| 13 | 1.358e-05 | 2.716e-05 | 1 |
| 14 | 2.196e-05 | 4.392e-05 | 1 |
| 15 | 0.0002575 | 0.0005149 | 0.9997 |
| 16 | 9.134e-15 | 1.827e-14 | 1 |
| 17 | 0.02029 | 0.04058 | 0.9797 |
| 18 | 7.934e-07 | 1.587e-06 | 1 |
| 19 | 0.0004561 | 0.0009122 | 0.9995 |
| 20 | 3.955e-16 | 7.91e-16 | 1 |
| 21 | 9.125e-15 | 1.825e-14 | 1 |
| 22 | 0.9971 | 0.00589 | 0.002945 |
| 23 | 7.077e-07 | 1.415e-06 | 1 |
| 24 | 0.6204 | 0.7591 | 0.3796 |
| 25 | 4.531e-12 | 9.062e-12 | 1 |
| 26 | 1.689e-14 | 3.377e-14 | 1 |
| 27 | 2.593e-10 | 5.185e-10 | 1 |
| 28 | 0.9996 | 0.0007369 | 0.0003684 |
| 29 | 0.9838 | 0.03232 | 0.01616 |
| 30 | 0.9909 | 0.01824 | 0.009121 |
| 31 | 0.4388 | 0.8776 | 0.5612 |
| 32 | 2.364e-05 | 4.728e-05 | 1 |
| 33 | 8.347e-07 | 1.669e-06 | 1 |
| 34 | 0.8224 | 0.3552 | 0.1776 |
| 35 | 0.0008974 | 0.001795 | 0.9991 |
| 36 | 3.69e-06 | 7.379e-06 | 1 |
| 37 | 1 | 1.083e-32 | 5.417e-33 |
| 38 | 0.8102 | 0.3797 | 0.1898 |
| 39 | 1 | 2.92e-14 | 1.46e-14 |
| 40 | 1.015e-08 | 2.03e-08 | 1 |
| 41 | 1 | 1.784e-14 | 8.921e-15 |
| 42 | 1.173e-17 | 2.345e-17 | 1 |
| 43 | 0.7183 | 0.5634 | 0.2817 |
| 44 | 3.929e-07 | 7.857e-07 | 1 |
| 45 | 0.3244 | 0.6488 | 0.6756 |
| 46 | 0.0008765 | 0.001753 | 0.9991 |
| 47 | 0.9998 | 0.0004244 | 0.0002122 |
| 48 | 0.9491 | 0.1018 | 0.05089 |
| 49 | 9.355e-36 | 1.871e-35 | 1 |
| 50 | 1 | 6.391e-13 | 3.196e-13 |
| 51 | 0.0003582 | 0.0007164 | 0.9996 |
| 52 | 0.6909 | 0.6183 | 0.3091 |
| 53 | 2.887e-12 | 5.774e-12 | 1 |
| 54 | 1.174e-16 | 2.349e-16 | 1 |
| 55 | 0.997 | 0.005922 | 0.002961 |
| 56 | 0.1985 | 0.3969 | 0.8015 |
| 57 | 0.04184 | 0.08367 | 0.9582 |
| 58 | 6.1e-16 | 1.22e-15 | 1 |
| 59 | 1 | 8.978e-14 | 4.489e-14 |
| 60 | 1 | 3.832e-09 | 1.916e-09 |
| 61 | 1 | 1.482e-11 | 7.41e-12 |
| 62 | 7.708e-35 | 1.542e-34 | 1 |
| 63 | 0.9825 | 0.03491 | 0.01746 |
| 64 | 0.009493 | 0.01899 | 0.9905 |
| 65 | 3.32e-06 | 6.64e-06 | 1 |
| 66 | 1.216e-05 | 2.433e-05 | 1 |
| 67 | 1 | 7.556e-07 | 3.778e-07 |
| 68 | 1 | 1.274e-11 | 6.368e-12 |
| 69 | 1 | 2.304e-05 | 1.152e-05 |
| 70 | 1 | 1.79e-12 | 8.95e-13 |
| 71 | 1.165e-38 | 2.331e-38 | 1 |
| 72 | 0.9588 | 0.08233 | 0.04116 |
| 73 | 0.005787 | 0.01157 | 0.9942 |
| 74 | 4.075e-12 | 8.15e-12 | 1 |
| 75 | 1 | 7.362e-07 | 3.681e-07 |
| 76 | 1 | 8.435e-16 | 4.218e-16 |
| 77 | 1 | 1.094e-20 | 5.472e-21 |
| 78 | 1 | 2.257e-05 | 1.128e-05 |
| 79 | 1 | 5.737e-05 | 2.868e-05 |
| 80 | 0.9932 | 0.01352 | 0.006758 |
| 81 | 0.3906 | 0.7812 | 0.6094 |
| 82 | 1 | 4.473e-19 | 2.237e-19 |
| 83 | 1 | 3.337e-13 | 1.669e-13 |
| 84 | 0.9981 | 0.003894 | 0.001947 |
| 85 | 1 | 1.045e-07 | 5.224e-08 |
| 86 | 1 | 8.097e-07 | 4.049e-07 |
| 87 | 1 | 4.704e-10 | 2.352e-10 |
| 88 | 1.753e-09 | 3.507e-09 | 1 |
| 89 | 1 | 5.423e-05 | 2.711e-05 |
| 90 | 1 | 6.25e-07 | 3.125e-07 |
| 91 | 0.9984 | 0.003164 | 0.001582 |
| 92 | 0.9976 | 0.004739 | 0.00237 |
| 93 | 1.74e-05 | 3.48e-05 | 1 |
| 94 | 0.2904 | 0.5808 | 0.7096 |
| 95 | 0.7586 | 0.4828 | 0.2414 |
| Meta Analysis of Goldfeld-Quandt test for Heteroskedasticity | |||
| Description | # significant tests | % significant tests | OK/NOK |
| 1% type I error level | 64 | 0.7273 | NOK |
| 5% type I error level | 71 | 0.806818 | NOK |
| 10% type I error level | 73 | 0.829545 | NOK |
| Ramsey RESET F-Test for powers (2 and 3) of fitted values |
> reset_test_fitted RESET test data: mylm RESET = 0.30187, df1 = 2, df2 = 96, p-value = 0.7401 |
| Ramsey RESET F-Test for powers (2 and 3) of regressors |
> reset_test_regressors RESET test data: mylm RESET = 1.735, df1 = 8, df2 = 90, p-value = 0.101 |
| Ramsey RESET F-Test for powers (2 and 3) of principal components |
> reset_test_principal_components RESET test data: mylm RESET = 1.166, df1 = 2, df2 = 96, p-value = 0.316 |
| Variance Inflation Factors (Multicollinearity) |
> vif TVDC2 TVDC3 TVDC4 TVDCSUM 1.896117 2.867499 1.408976 4.888353 |